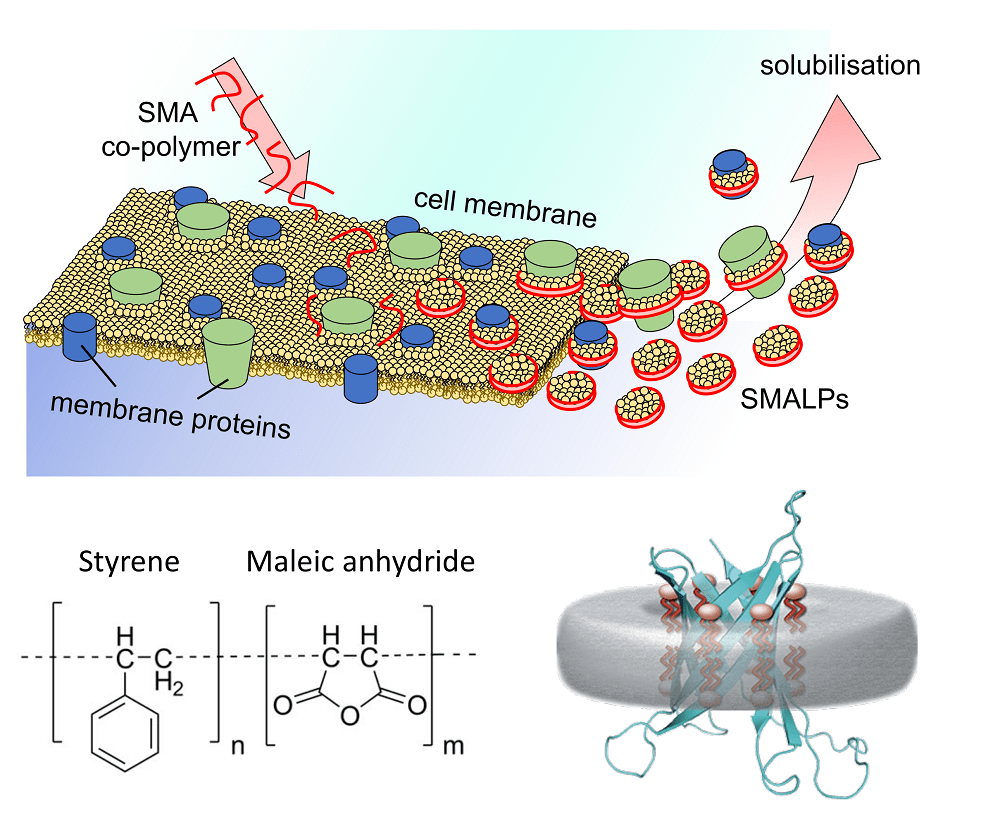
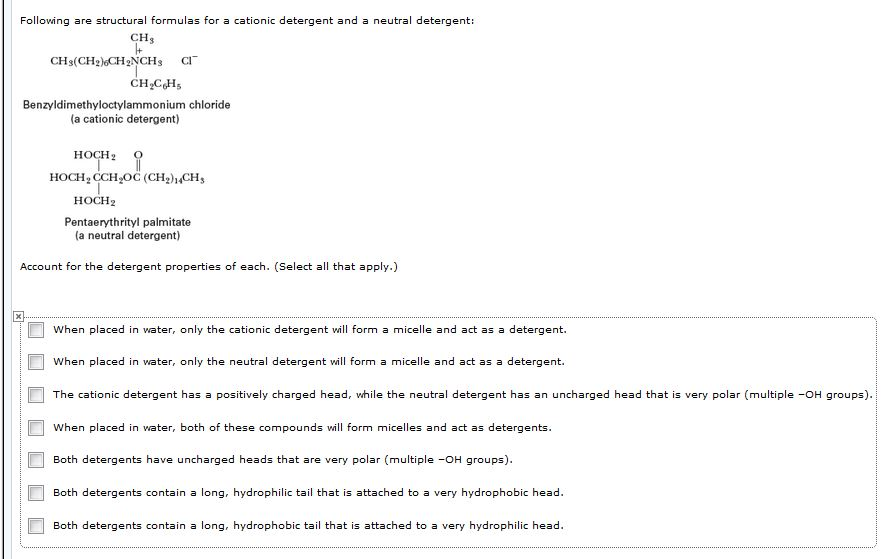
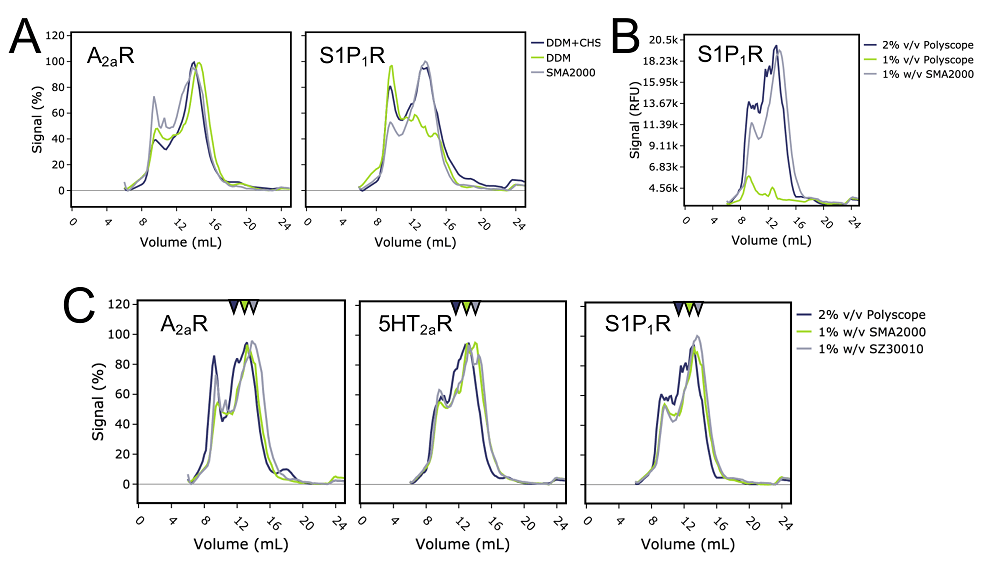
Detergents: Friends not foes for high‐performance membrane proteomics toward precision medicine - Zhang - 2017 - PROTEOMICS - Wiley Online Library

Chemical structures of DPPC, CHOL, CHS, and CHSA molecules with the... | Download Scientific Diagram
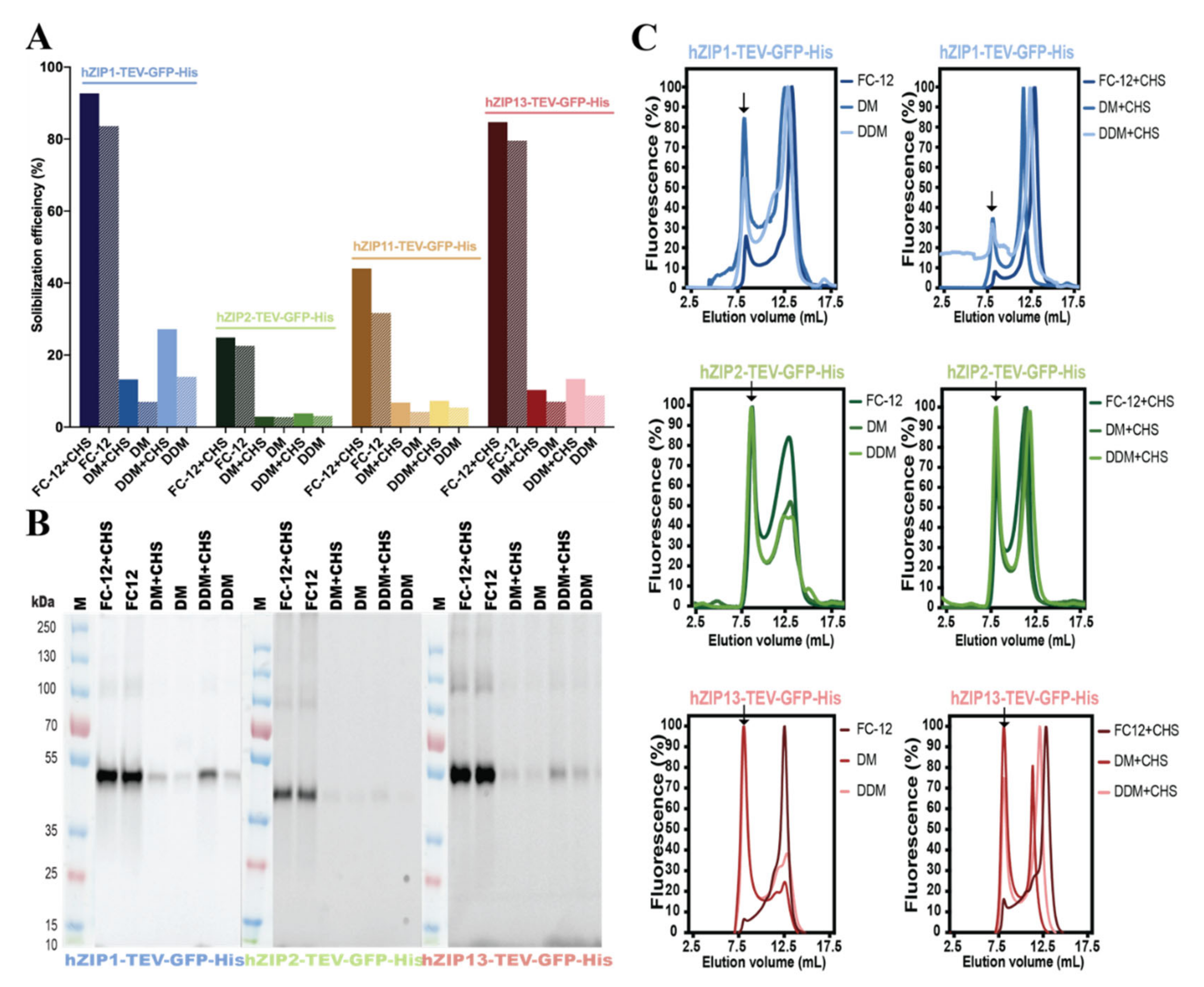
Cells | Free Full-Text | Overproduction of Human Zip (SLC39) Zinc Transporters in Saccharomyces cerevisiae for Biophysical Characterization
Cryo-EM structure of the mitochondrial protein-import channel TOM complex from Saccharomyces cerevisiae
Sequential purification and characterization of Torpedo californica nAChR-DC supplemented with CHS for high-resolution crystalli

Tricks of the trade used to accelerate high-resolution structure determination of membrane proteins - ScienceDirect

Quantitative analysis of sterol-modulated monomer–dimer equilibrium of the β1-adrenergic receptor by DEER spectroscopy | PNAS
Stabilization of Functional Recombinant Cannabinoid Receptor CB2 in Detergent Micelles and Lipid Bilayers | PLOS ONE

Experimental determination and computational interpretation of biophysical properties of lipid bilayers enriched by cholesteryl hemisuccinate - ScienceDirect

RCSB PDB - 6F3W: Backbone structure of free bradykinin (BK) in DDM/CHS detergent micelle determined by MAS SSNMR
.png)